 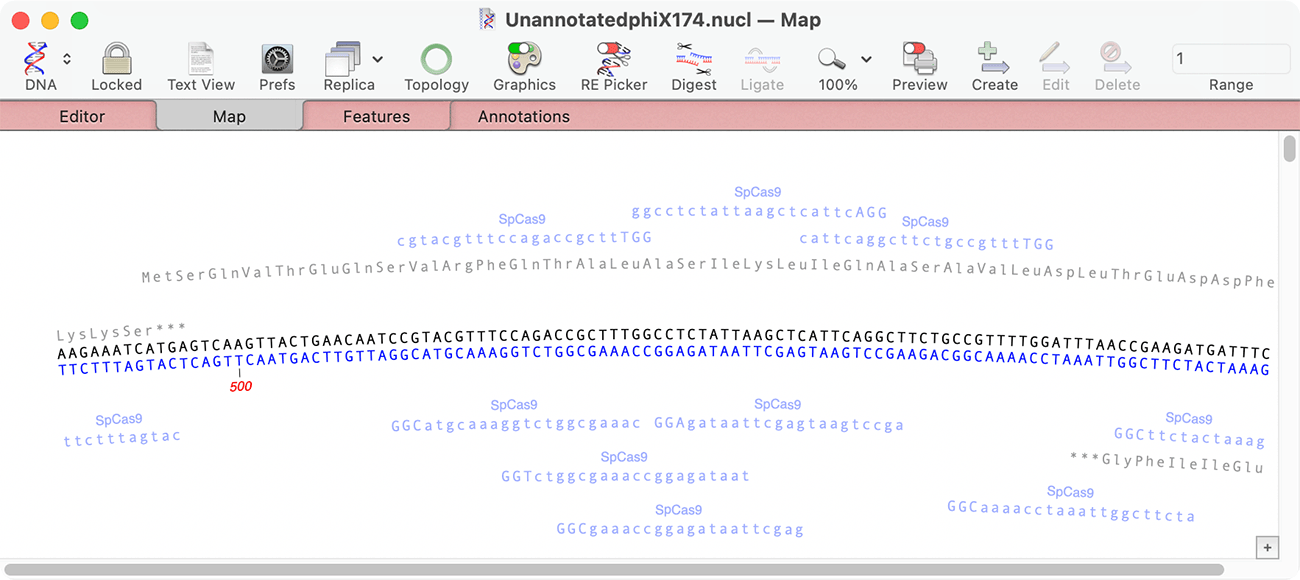
If you wish to compare two or more DNA sequences, you should definitely consider if one of the other alignment functions may be more suitable. This functionality is most suited for protein alignments, or for nucleic acid sequences where you are interested in examining phylogenetic relationships. Click on the Prefs toolbar button to control the appearance and behavior of the data in each of tabs that represent different views or analyses of the alignment. Add sequences to the alignment by using the Edit->Add Sequences from File menu item then click on the Align toolbar button to automatically align the sequences using ClustalW. Choose File->New->Protein Alignment (or File->New->Nucleic Acid Alignment) to create an empty MSA window. If you have two or more related sequences (DNA or Protein) and you want to examine the relationship between them, use this function. Pustell Matrix (also known as a Dot Plot)ĬlustalW/Multiple Sequence Alignment (MSA).ClustalW – we also call this the “standard” Multiple Sequence Alignment (MSA).Each function is designed for a different purpose. They say there’s more than one way to skin a cat (not that I’ve done that – I have skinned a catfish, but I only know one way), and thats certainly true for alignments in MacVector. We get a lot of comments and questions from users on the various alignment functions in MacVector. If you already own an older copy of MacVector, and would like to know what new features have been added since the version you own, you can find that information on this summary page.Update 19 August 2013: We’ve added support for Muscle and T-Coffee to the MSA editor MacVector can read and write DNA and Protein sequences in most popular file formats. In addition to directly editing sequences and features/annotations, MacVector has an intuitive "Click Cloning" graphical interface that lets you easily replicate laboratory cloning experiments to create new molecules. MacVector uses the native Mac OS X Quartz graphics to generate publication quality images that can be scaled to any size with no loss of resolution. MacVector supports the popular Gateway, TOPO TA and Zero Blunt cloning technologies from Invitrogen. With a few simple clicks from an intuitive graphical interface, you can replicate your biological manipulations at the bench to create new molecules with the correct sequences across the cloning and recombination junctions. You can scan an unannotated or partially annotated sequence against a folder on your hard drive and MacVector will identify matching features in sequences in the target folder and add them to your sequence. Because MacVector includes custom feature appearance information when annotating the sequence, you can use this to maintain a carefully curated set of your favorite genes and sequences each with a graphical appearance that best suits your needs. You can design primers for either PCR or Sequencing/Hybridization probes using a variety of primer design functions, including an interactive manual design interface along with several scanning functions such as the popular Primer3 algorithm. The interactive interface clearly shows which potential primers may have secondary structure problems, alternate binding sites or other characteristics that might impact their use in experiments. MacVector provides a wide variety of useful DNA analysis tools, including base composition analysis, Restriction Enzyme searches, DNA Subsequence searches and "Dot-Plot" comparisons between DNA:DNA and DNA:Protein sequences. A Coding Preference toolbox lets you select a variety of algorithms to graphically scan a DNA sequence for likely protein coding open reading frames. Protein sequences can be reverse translated into DNA, compared using "Dot-Plot" analysis and scanned for Proteolytic cleavage sites and amino acid sequence motifs. A comprehensive Protein Analysis Toolbox provides a wide variety of algorithms for analyzing the composition of proteins and presenting the results in graphical and tabular formats. MacVector has built-in Internet connectivity to the NCBI BLAST and Entrez databases. You can directly search Entrez for DNA or Protein sequences based on features, authors, keywords etc and directly download them into MacVector, complete with all features and annotations. 
The built-in BLAST interface lets you submit multiple BLAST jobs using DNA or Protein sequences and then download any matching sequences by selecting them from a hit list. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
